What a journey — I’ve been working on this for a couple of months now and I am proud to present the result. Did you ever get into a position where you might have multiple ideas for analyzing a specific field or domain but just lack the data? It is a tedious process… cleaning, preprocessing, architecture, reliability, availability… the list goes on. So I asked myself: wouldn’t it be nice to have a single source of data for my science-of-science projects? One sort of data lake to handle everything I would throw at it? Citation analyses, disruption and impact indexing, concept mining and taxonomy generation. This is when the Science Data Lake project idea was born.
The paper is now available on arXiv (2603.03126), the dataset lives on HuggingFace, and all code is on GitHub.
The problem: scholarly data is a mess
If you have ever tried to do large-scale bibliometric research, you know the pain. Semantic Scholar has citation counts and open-access flags. OpenAlex offers field-weighted citation impact, topics, and funding data. SciSciNet provides disruption indices and atypicality scores. Papers with Code links papers to repositories. Retraction Watch tracks integrity events. Reliance on Science maps patent-to-paper citations. Each source is individually excellent — but no single database captures all facets simultaneously.
Want to study whether papers that release code are more or less disruptive? You need Papers with Code and SciSciNet and OpenAlex topic assignments. Want to check whether retracted papers show anomalous citation patterns across databases? You need Retraction Watch and at least two citation sources. Every researcher ends up writing their own ad-hoc integration scripts that are rarely shared, wasting effort and introducing inconsistencies.
The Science Data Lake
The Science Data Lake unifies eight open scholarly data sources into a single, locally-deployable infrastructure. At its core, it is a collection of Apache Parquet files (about 960 GB total) that are exposed through a lightweight DuckDB database (~270 KB) defining 153 SQL views across 22 schemas.
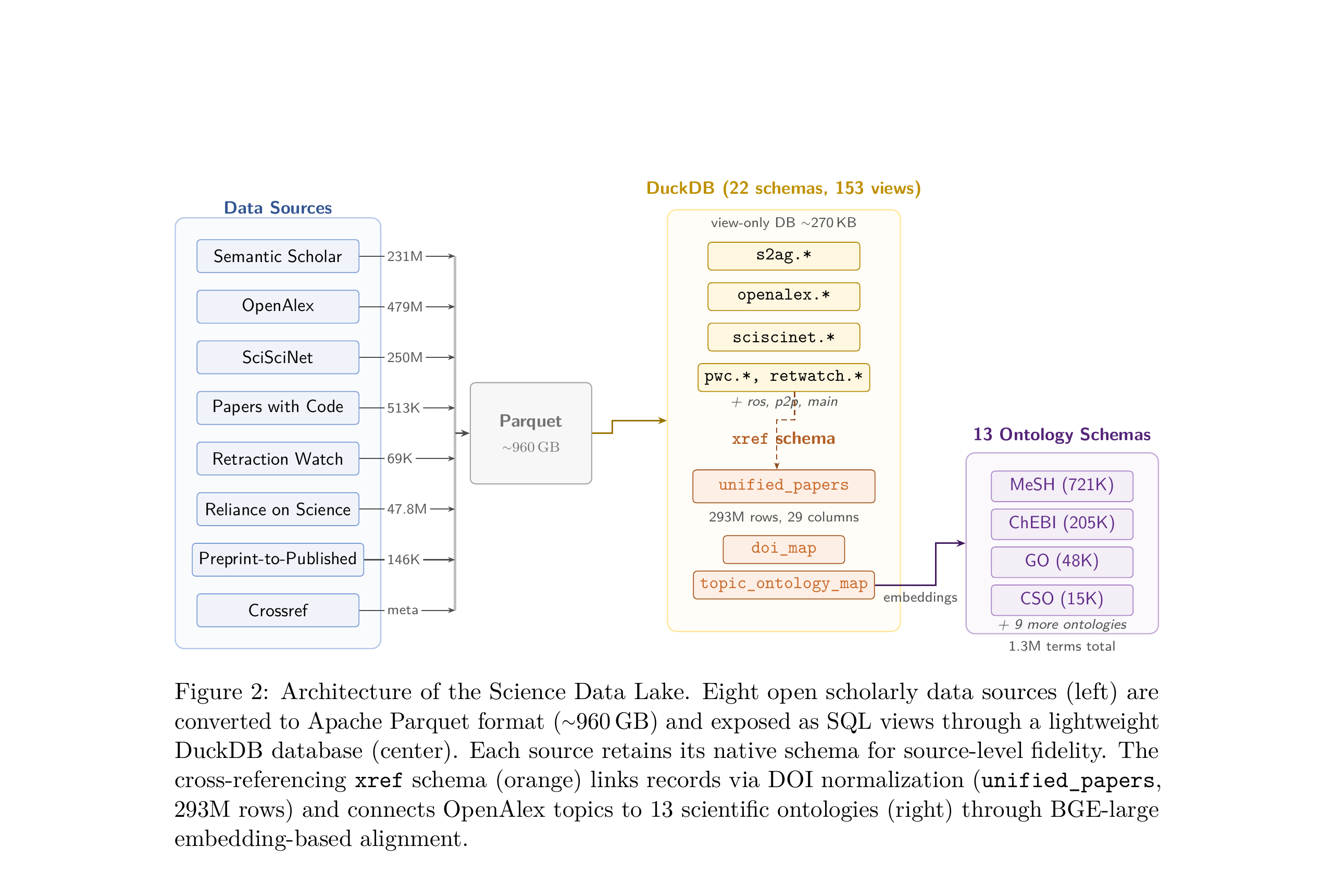 Eight data sources are converted to Parquet and exposed as SQL views through DuckDB. The cross-referencing xref schema links records via DOI normalization and connects topics to 13 scientific ontologies.
Eight data sources are converted to Parquet and exposed as SQL views through DuckDB. The cross-referencing xref schema links records via DOI normalization and connects topics to 13 scientific ontologies.
The design follows two key principles:
Source-level preservation. Each data source retains its native schema within a dedicated namespace — s2ag.papers, openalex.works, sciscinet.paper_metrics, and so on. You can directly inspect how different databases describe the same paper without any information loss from schema merging.
Cross-referencing via DOI normalization. Three materialized views bridge across sources: doi_map handles DOI normalization (different sources store DOIs in annoyingly different formats), unified_papers creates a 293-million-row join table with 29 columns spanning all sources, and topic_ontology_map connects OpenAlex topics to formal scientific ontologies.
What’s inside
| Source | Records | What it adds |
|---|---|---|
| Semantic Scholar (S2AG) | 231M | Citations, influential citations, open access |
| OpenAlex | 479M | FWCI, topics, types, languages, funding |
| SciSciNet | 250M | Disruption, atypicality, team size |
| Papers with Code | 513K | Code repositories, tasks, datasets |
| Retraction Watch | 69K | Retraction reasons and dates |
| Reliance on Science | 47.8M | Patent-paper citation pairs |
| Preprint-to-Published | 146K | bioRxiv/medRxiv DOI mappings |
| Crossref | — | DOI metadata, reference lists |
The unified_papers table lets you query across all of these simultaneously with a single SQL statement — something that would require juggling multiple APIs and custom join logic otherwise.
Bridging vocabularies: embedding-based ontology alignment
One of the more interesting technical challenges was connecting OpenAlex’s topic taxonomy to formal scientific ontologies. OpenAlex assigns papers to 4,516 topics organized in four hierarchical levels (domain, field, subfield, topic), but these topics don’t map to established ontologies like MeSH (medicine), Gene Ontology (biology), or the Computer Science Ontology.
I developed an embedding-based alignment method: compute BGE-large sentence embeddings for both OpenAlex topics and ontology terms (1.3 million terms across 13 ontologies), then find nearest neighbors using a FAISS index. This yields 16,150 mappings covering 99.8% of all topics.
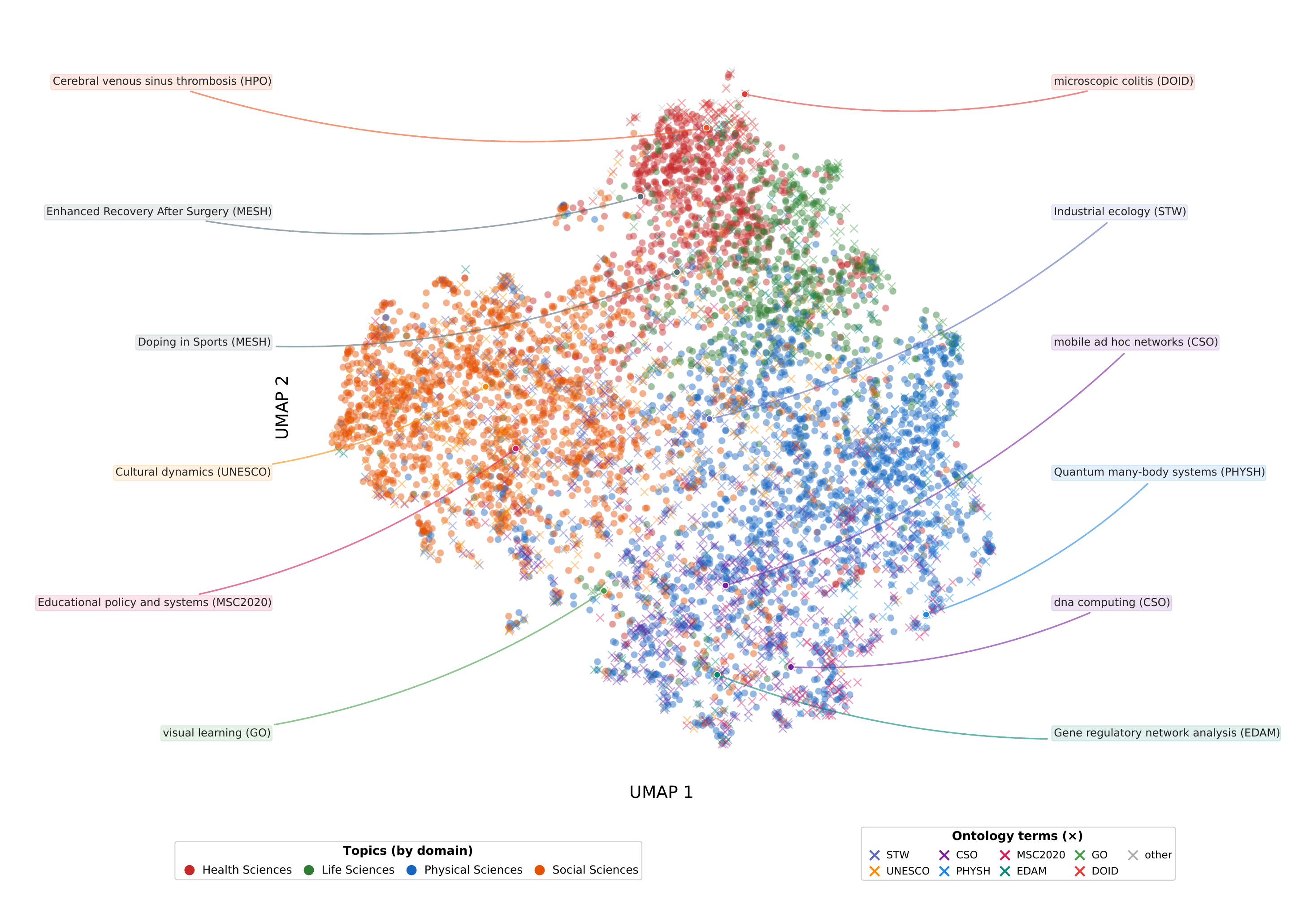 UMAP projection of BGE-large embeddings for OpenAlex topics (points, colored by domain) and matched ontology terms (crosses). Semantic clusters emerge naturally — health science topics co-locate with MeSH terms, computer science topics with CSO terms, and so on.
UMAP projection of BGE-large embeddings for OpenAlex topics (points, colored by domain) and matched ontology terms (crosses). Semantic clusters emerge naturally — health science topics co-locate with MeSH terms, computer science topics with CSO terms, and so on.
At the recommended similarity threshold (), the method achieves , outperforming TF-IDF cosine (), BM25 (), and Jaro-Winkler () on a 300-pair gold-standard evaluation. The key advantage is capturing semantic similarity that lexical methods miss — for example, “Artificial Intelligence in Medicine” maps to EDAM’s “Medical informatics” (cosine similarity 0.87), a connection no string-matching method would find.
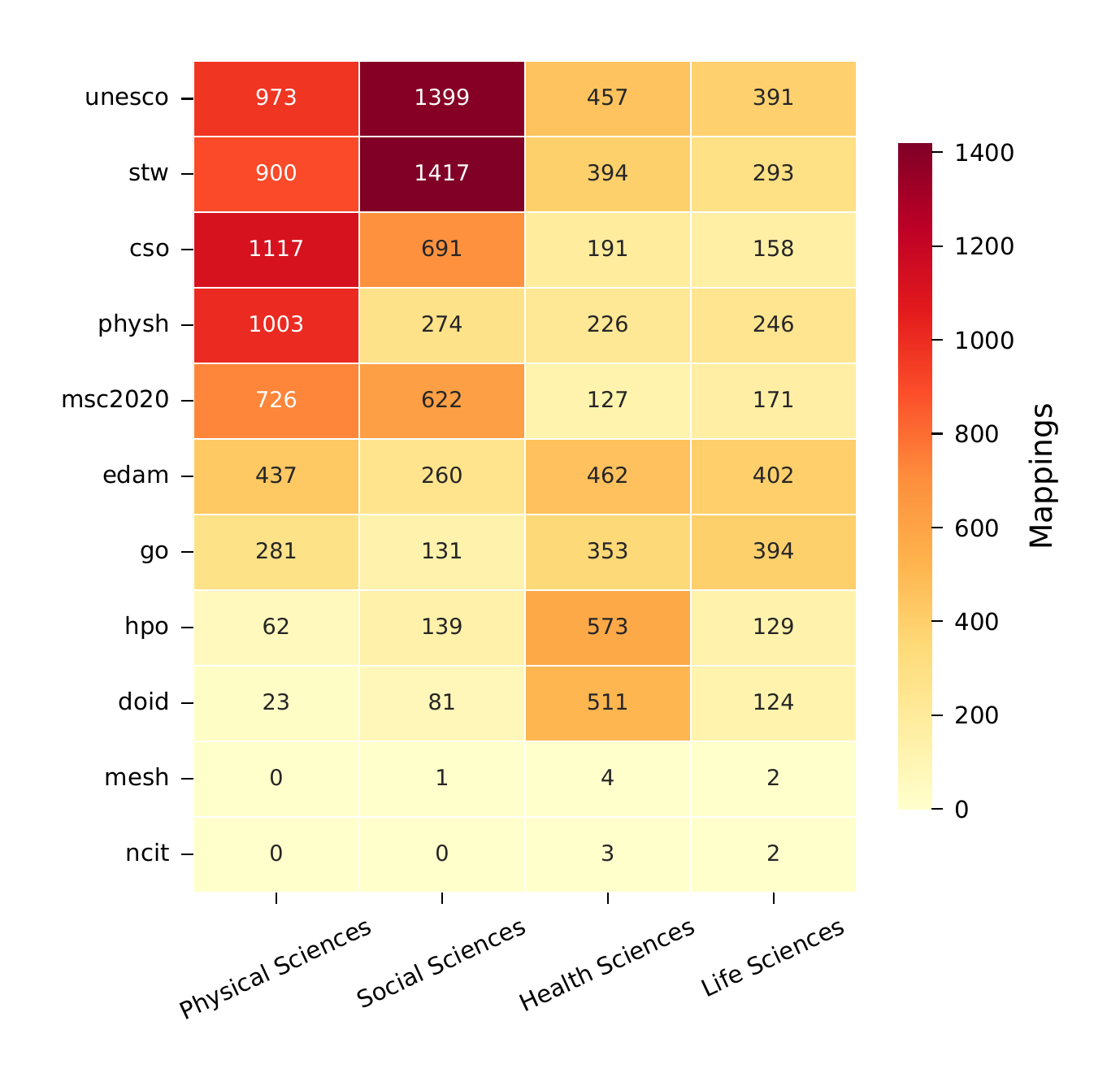 Heatmap showing high-quality ontology mappings by OpenAlex domain. Different ontologies specialize in different scientific areas: MeSH dominates health sciences, CSO covers computer science, GO spans molecular biology, and AGROVOC bridges agricultural and environmental sciences.
Heatmap showing high-quality ontology mappings by OpenAlex domain. Different ontologies specialize in different scientific areas: MeSH dominates health sciences, CSO covers computer science, GO spans molecular biology, and AGROVOC bridges agricultural and environmental sciences.
Validation: does it actually work?
I validated the data lake through three approaches:
10 automated sanity checks — all pass without violations. These verify DOI format consistency, coverage flag accuracy, primary key uniqueness, ontology map integrity, and more.
Cross-source citation agreement — for the ~121 million papers present in all three major citation sources (S2AG, OpenAlex, SciSciNet), pairwise Pearson correlations range from to . The sources broadly agree, but the non-negligible differences (particularly S2AG vs. OpenAlex at ) underscore why preserving all three independent counts matters for sensitivity analyses.
Manual ontology annotation — a stratified sample of 300 topic-ontology pairs was labeled as correct, partial, or incorrect. Zero incorrect mappings appeared in the exact and high-quality tiers.
Four things you can only do with the Data Lake
To demonstrate the value of multi-source integration, the paper presents four vignettes — each requiring data from multiple sources that no single database provides:
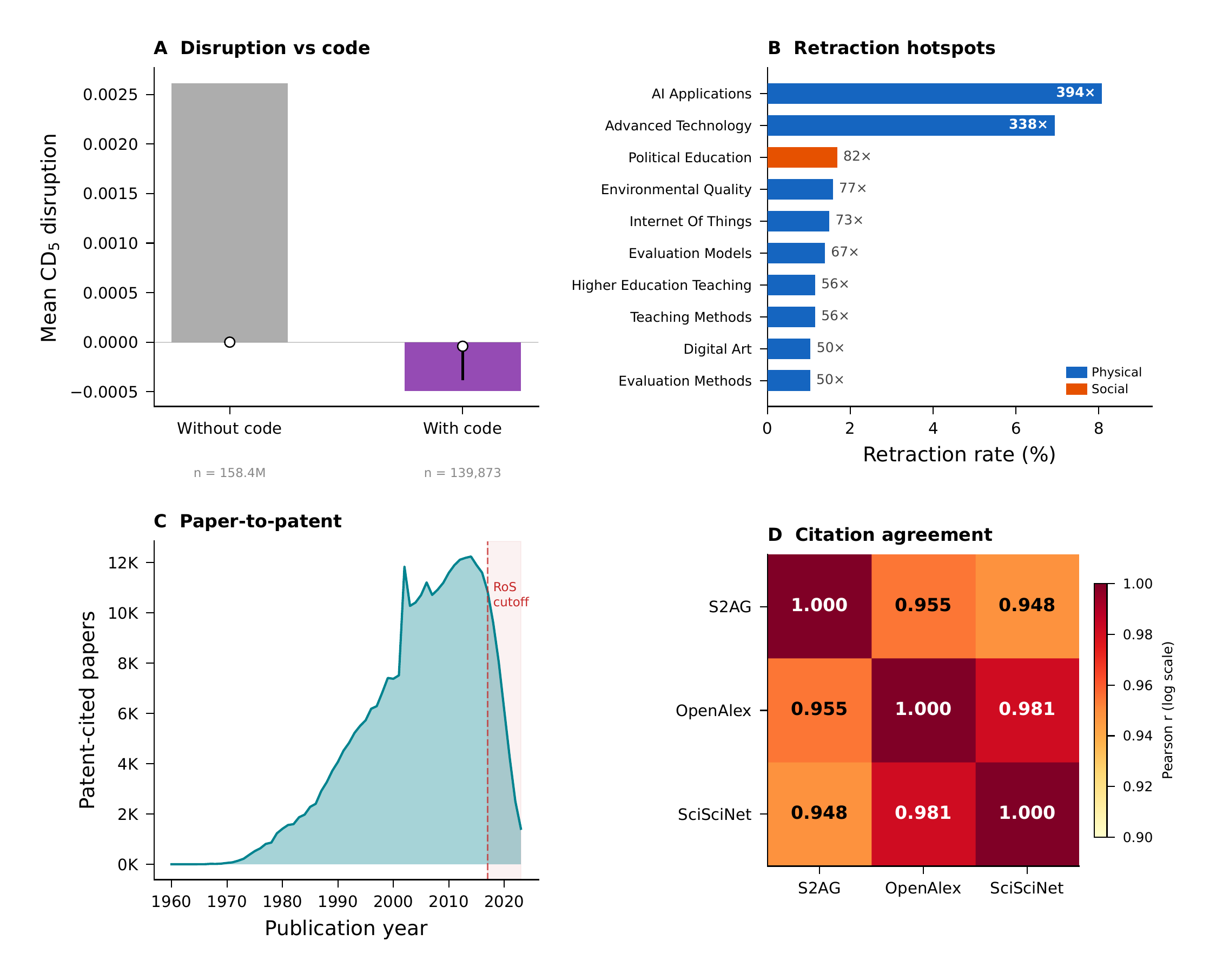 Four vignettes showing analyses that are only possible through cross-source integration.
Four vignettes showing analyses that are only possible through cross-source integration.
1. Disruption vs. code availability. Papers with code repositories show a mean disruption () of , compared to for papers without code. Code-releasing papers tend to be slightly more consolidating (building on existing work) rather than disruptive. This varies across ontology domains — computer science topics show the strongest code adoption, while biomedical topics show lower code rates but higher disruption variability. This analysis requires SciSciNet (disruption), Papers with Code (code flags), OpenAlex (topics), and ontology bridging — four resources from four different sources.
2. Retraction enrichment by domain. Retracted papers show a mean disruption of 0.0035 compared to 0.0026 for non-retracted papers. More strikingly, ontology-level enrichment analysis reveals retraction hotspots: topics mapped to “AI Applications” show enrichment, and “Advanced Technology” topics show enrichment relative to baseline retraction rates.
3. Patent-citation impact. Patent-cited papers have dramatically higher impact: mean citation count of 94.3 versus 16.1 for non-patent-cited papers (), and mean FWCI of 4.7 versus 1.5 (). The multi-ontology footprint reveals that patent-cited papers cluster in MeSH-mapped health sciences, CSO-mapped computer science, and ChEBI-mapped chemistry topics.
4. Cross-source citation reliability. Restricting to 121 million papers present in all three sources, pairwise citation correlations range from 0.76 to 0.87. The mean absolute differences range from 2.3 to 4.1 citations. The most extreme outlier: a paper with 257,887 citations in S2AG and zero in OpenAlex — showing how coverage differences can produce dramatic record-level discrepancies.
Built for AI-assisted research
One design choice I’m particularly excited about is the structured schema reference (SCHEMA.md, ~1,200 lines). It documents every table name, column, data type, row count, and cross-dataset join strategy in a format optimized for LLM consumption. A query like “find the most disruptive papers in computer science that have open-source code and check their retraction status” requires joining four schemas (sciscinet, xref, pwc, retwatch) — and SCHEMA.md gives an LLM agent all the metadata it needs to formulate those joins without additional context.
How to get started
The entire infrastructure is open source:
- Dataset: HuggingFace (DOI: 10.57967/hf/7850)
- Code: GitHub
- Paper: arXiv 2603.03126
For local deployment, clone the repo and run datalake_cli.py — it downloads, converts, and links all sources automatically. For quick exploration without downloading ~1 TB of data, you can query HuggingFace-hosted Parquet files directly through DuckDB’s httpfs extension. All you need is DuckDB and a SQL prompt.
What’s next
The architecture is designed to be extensible — adding a new data source means writing a Parquet converter and registering the schema. I’m looking forward to seeing what the research community does with this. Whether it’s large-scale bibliometric studies, science policy analysis, or feeding structured scholarly data to AI research agents — the Science Data Lake provides the foundation.
If you have questions, ideas, or want to contribute additional sources or ontologies, feel free to reach out or open an issue on GitHub.
